Overview
The group's research lines fall within the field of bacterial pathogenesis and antimicrobial resistance (PatoBAnt). Antibiotic resistance is one of the most challenging problems in medicine and biology, and in particular, healthcare-associated infections caused by multidrug-resistant (MDR) bacteria are becoming a main cause of public health concern. Our group has been working at the IBB since 2000 and our main activity is the development of new strategies to try to solve this problem of marked socio-sanitary character. Thus, our research have been focused on the study of virulence and resistance determinants in different human pathogens such as Stenotrophomonas maltophilia, Pseudomonas aeruginosa, Burkholderia sp., Acinetobacter baumannii, Staphylococcus aureus and Mycobacterium tuberculosis among others. We try to unveil the genomic and molecular bases, from a genetic and functional point of view, of processes involved in pathogenesis, virulence and drug resistance. In recent years we have focused much of our efforts on S. maltophilia, as a model of an emerging nosocomial MDR microorganism. Thus, we aim to deepen the functional knowledge of its pathogenesis processes and exploit this knowledge to identify new targets for the development of novel drugs or alternative strategies to current treatments. In particular, we focus on the study of non-essential physiological processes such as the quorum sensing system and regulatory elements of gene expression as intrinsic determinants of virulence and resistance. The design of drugs against non-essential processes circumvents the selection of resistant strains and is therefore expected to result in a decrease in resistance development. In collaboration with groups at IBB and other outside groups, we apply a multidisciplinary approach, ranging from classical microbiology and bacterial molecular genetics to genomics, proteomics, and bioinformatics that should end with a selection of validated targets and drugs to suppress bacterial virulence and/or resistance phenotypes.
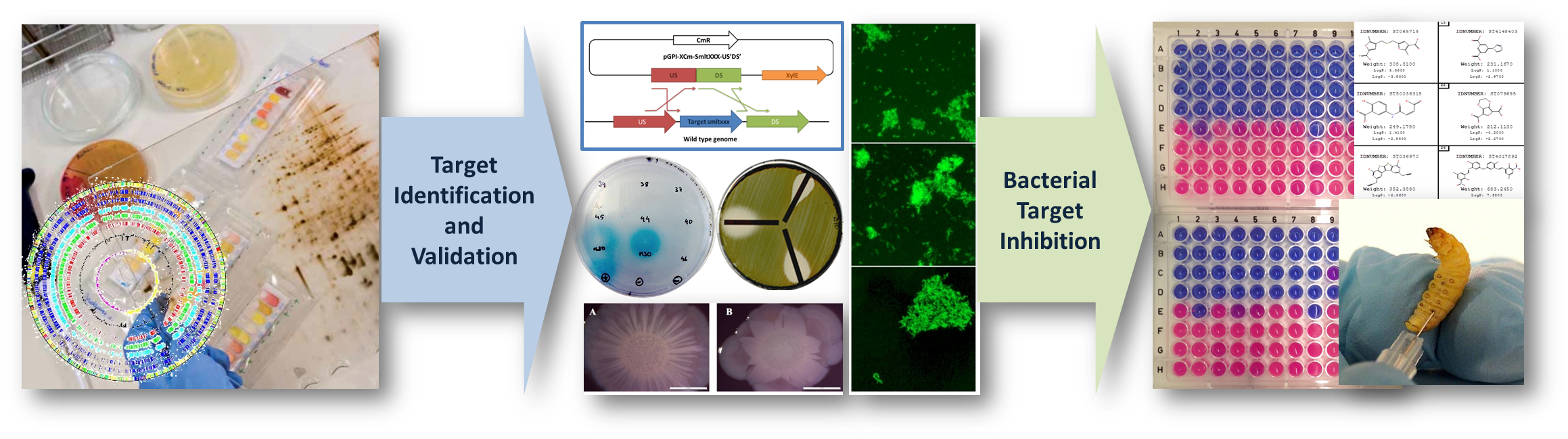
Currently we have four research lines:
Molecular and genomic bases of bacterial pathogenesis and antimicrobial agent resistance.
This project concerns the search and characterization of targets involved in the modulation of virulence and resistance in MDR Gram-negative pathogens. To address this general work plan we combine methods from microbial genetics, comparative genomics, transcriptomic analysis, structural biology and bioinformatics within a pan-genomic strategy. The main objectives that are addressed are: i) selection of "type strains" based on phenotypic and genotypic data and subsequent sequencing and annotation of their genomes when not available; ii) comparative genomics and determination of shared proteomes and exclusive proteomes of the most extreme strains in terms of virulence and/or drug resistance (in collaboration with the Computational Biology group led by Xavier Daura); iii) molecular epidemiology and genotype-phenotype correlations; iv) study of intrinsic physiological mechanisms involved in virulence/resistance by generation of specific mutants and evaluation of their phenotypes and virulence in different alternative animal models (Caenorhabditis elegans and Galleria mellonella, and zebrafish in collaboration with the Evolutive Immunology group led by Nerea Roher); and v) final selection and validation of candidate targets on the basis of the previous results.
Quorum sensing signal-response systems in Stenotrophomonas maltophilia.
In nosocomial settings, although with moderate incidence, S. maltophilia can cause severe infections, mainly in immunocompromised patients. Once in a human host, S. maltophila is difficult to eradicate due to its ability to form biofilm and its natural antibiotic resistance arsenal. Cell-to-cell signalling processes in S. maltophilia, controlled by the quorum-sensing (QS) systems, are thought to be pivotal in the regulation of several of these virulence and resistance factors. Our group aims at deepening the understanding of the QS networks of S. maltophilia at functional and molecular levels and exploiting this knowledge for the identification of targets for the development of an antimicrobial strategy based on the quenching of quorum sensing. Antimicrobial compounds against these targets could be potentially used to reduce the pathogenic capacity of the bacteria and/or potentiate the activity of current antibiotics.
Study of the mechanisms of resistance to colistin in Gram-negative bacteria.
One of the antibiotics the group has focused on in recent years has been colistin. Colistin (polymyxin E) is an antibiotic used as a last-resort for MDR Gram-negative pathogens. Colistin is a polycationic peptide that interacts with the bacterial lipopolysaccharide (LPS) in the outer membrane and through its hydrophobic portion disrupts bacterial cell membranes causing cell lysis. The emergence of colistin-resistant isolates, as for other antibiotics, is considered a serious problem that requires special attention as it is a drug of restricted use. Our group has recently alerted about the existence of adaptive resistance and heteroresistance to colistin in S. maltophilia. We are currently identifying the genetic determinants of colistin resistance and applying genomic sequencing and transcriptome-based analyses to reveal the mechanisms governing these phenomena of heterogeneous resistance. In addition, and in collaboration with Uwe Mamat´s and Nicolas Gisch´s labs at the Research Center Borstel (RCB, Germany), we are studying LPS modifications as a mechanism of colistin resistance in S. maltophilia.
Validation of novel hit compounds as potential antibacterial drugs against Gram-negative pathogens.
In collaboration with the Computational Biology group at IBB led by Xavier Daura, we work on the design of antibacterial hits identified by virtual screening of chemical compounds against key targets of the pathogenesis and virulence processes. The drug discovery process and experimental validation of hit compounds includes: i) target proposal based on experimental evidences; ii) analysis of the intrinsic antimicrobial activity of selected compounds against and extended panel of MDR Gram-negative organisms; iii) enhancer activity of the hit compounds in combination with known antibiotics; iv) determination of the inhibitory effect of the compounds on biofilm formation; v) efficacy assays based on the analysis of antimicrobial or antivirulence activity in vivo in different animal models; and vi) in vitro assessment of the induction of drug resistance. Furthermore, we are evaluating, in collaboration with Timothy O'Sullivan at University College Cork (Ireland), molecules that interfere with the bacteria's native QS communication system and prevent them from producing biofilm.