https://doi.org/10.1016/j.jmb.2021.167059
Highlights
In this review, we discuss the applications and limitations of AlphaFold in the field of protein aggregation.•
AlphaFold might help in the computationally assisted optimization of the solubility of globular proteins with biomedical and industrial interest.•
In amyloid diseases, the heterogeneous nature of aggregation intermediates and amyloid fibrils hinders the use of AlphaFold.•
Residue covariation in functional amyloids suggests that AlphaFold could be trained to predict their structure, ultimately assisting in the design of amyloid-based nanomaterials.
Abstract
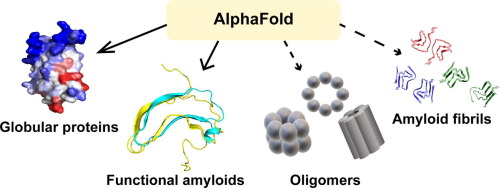
Protein aggregation is a widespread phenomenon with important implications in many scientific areas. Although amyloid formation is typically considered as detrimental, functional amyloids that perform physiological roles have been identified in all kingdoms of life. Despite their functional and pathological relevance, the structural details of the majority of molecular species involved in the amyloidogenic process remains elusive. Here, we explore the application of AlphaFold, a highly accurate protein structure predictor, in the field of protein aggregation. While we envision a straightforward application of AlphaFold in assisting the design of globular proteins with improved solubility for biomedical and industrial purposes, the use of this algorithm for predicting the structure of aggregated species seems far from trivial. First, in amyloid diseases, the presence of multiple amyloid polymorphs and the heterogeneity of aggregation intermediates challenges the “one sequence, one structure” paradigm, inherent to sequence-based predictions. Second, aberrant aggregation is not the subject of positive selective pressure, precluding the use of evolutionary-based approaches, which are the core of the AlphaFold pipeline. Instead, amyloid polymorphism seems to be constrained by the need for a defined structure-activity relationship in functional amyloids. They may thus provide a starting point for the application of AlphaFold in the amyloid landscape.
