Int. J. Mol. Sci. 2020, 21(16), 5814; https://doi.org/10.3390/ijms21165814
Santos, J.; Iglesias, V.; Pintado, C.; Santos-Suárez, J.; Ventura, S. DispHred: A Server to Predict pH-Dependent Order–Disorder Transitions in Intrinsically Disordered Proteins. Int. J. Mol. Sci. 2020, 21, 5814.
Abstract
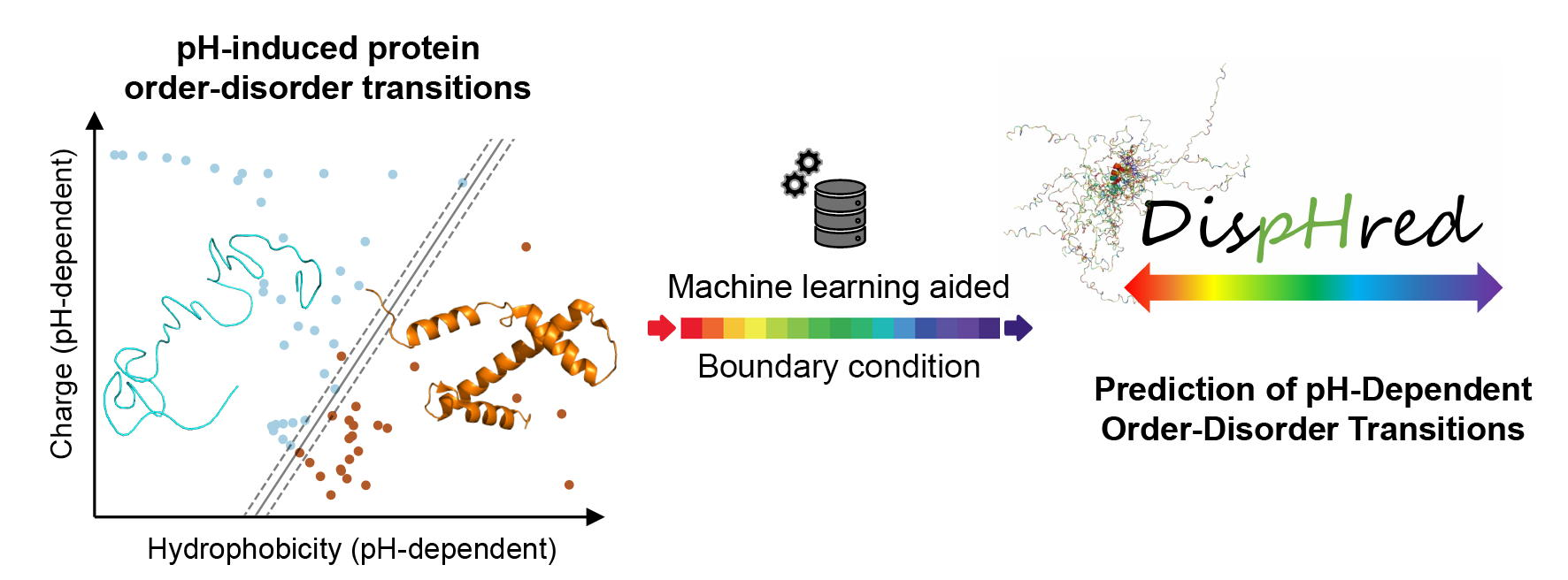
The natively unfolded nature of intrinsically disordered proteins (IDPs) relies on several physicochemical principles, of which the balance between a low sequence hydrophobicity and a high net charge appears to be critical. Under this premise, it is well-known that disordered proteins populate a defined region of the charge–hydropathy (C–H) space and that a linear boundary condition is sufficient to distinguish between folded and disordered proteins, an approach widely applied for the prediction of protein disorder. Nevertheless, it is evident that the C–H relation of a protein is not unalterable but can be modulated by factors extrinsic to its sequence. Here, we applied a C–H-based analysis to develop a computational approach that evaluates sequence disorder as a function of pH, assuming that both protein net charge and hydrophobicity are dependent on pH solution. On that basis, we developed DispHred, the first pH-dependent predictor of protein disorder. Despite its simplicity, DispHred displays very high accuracy in identifying pH-induced order/disorder protein transitions. DispHred might be useful for diverse applications, from the analysis of conditionally disordered segments to the synthetic design of disorder tags for biotechnological applications. Importantly, since many disorder predictors use hydrophobicity as an input, the here developed framework can be implemented in other state-of-the-art algorithms. View Full-Text
