 Abstract
Abstract
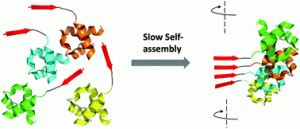
Amyloids have been exploited to build amazing bioactive materials. In most cases, short synthetic peptides constitute the functional components of such materials. The controlled assembly of globular proteins into active amyloid nanofibrils is still challenging, because the formation of amyloids implies a conformational conversion towards a β-sheet-rich structure, with a concomitant loss of the native fold and the inactivation of the protein. There is, however, a remarkable exception to this rule: yeast prions. They are singular proteins able to switch between a soluble and an amyloid state. In both states, the structure of their globular domains remains essentially intact. The transit between these two conformations is encoded in prion domains (PrDs): long and disordered sequences to which the active globular domains are appended. PrDs are much larger than typical self-assembling peptides. This seriously limits their use for nanotechnological applications. We have recently shown that these domains contain soft amyloid cores (SACs) that suffice to nucleate their self-assembly reaction. Here we genetically fused a model SAC with different globular proteins.
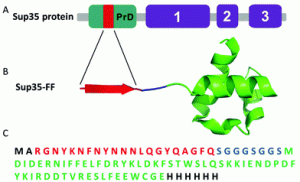 We demonstrate that this very short sequence acts as a minimalist PrD, driving the selective and slow assembly of the initially soluble fusion proteins into amyloid fibrils in which the globular proteins retain their native structure and display high activity. Overall, we provide here a novel, modular and straightforward strategy to build active protein-based nanomaterials at a preparative scale.
We demonstrate that this very short sequence acts as a minimalist PrD, driving the selective and slow assembly of the initially soluble fusion proteins into amyloid fibrils in which the globular proteins retain their native structure and display high activity. Overall, we provide here a novel, modular and straightforward strategy to build active protein-based nanomaterials at a preparative scale.
https://pubs.rsc.org/en/content/articlelanding/2019/nr/c9nr01755k#!divAbstract
